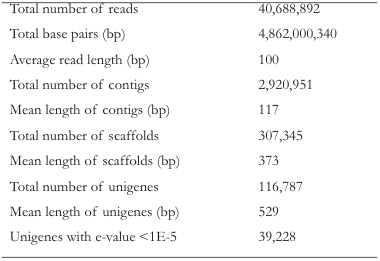
From Discussions VOL. 10 NO. 1Genome Profile of Ambystoma mexicanumRESULTS AND DISCUSSIONIllumina NGS and Read Assembly cDNA samples from Illumina sequencing were compared to the denervated limb stump as the control. Cumulatively, both libraries yielded over 40 million reads. Of the three de novo assemblies, SOAPdenovo produced the highest number of high-quality contigs at 2,920,951 contigs, whereas Trinity produced a smaller number of contigs but at a higher average read length. Due to its more prominent as the standard for the experiment (Li et al., 2010). From the SOAPdenovo data, unigenes, scaffolds that do not match any other scaffolds in the Xenopus levis database, numbered at 116,787, with a mean length of 529bp (Table 1) . In comparison, Monaghan and his colleagues used pyrosequencing to generate over 1,700,000 reads with approximately 400,000 unigenes from cDNA samples across a broad range of regeneration stages (Monaghan et al., 2009). The lower unigene count found in this experiment can be attributed to the small sample size used: tissue samples were harvested only once from each of the only 36 axolotls. Also, the average length of contigs synthesized by pyrosequencing is 215bp compared to 117bp by Illumina (Table 1), and because the longer the contigs, the more likely it is for the assembled scaffold to have no pre-existing matches and be unigene count (Tamura et al., 2008). Nevertheless, the fact that significantly more DNA was sequence by Illumina highlights its potential in analyzing genomic sequences. Table 1. SOAPdenovo analysis of the Ambystoma mexicanum cDNA
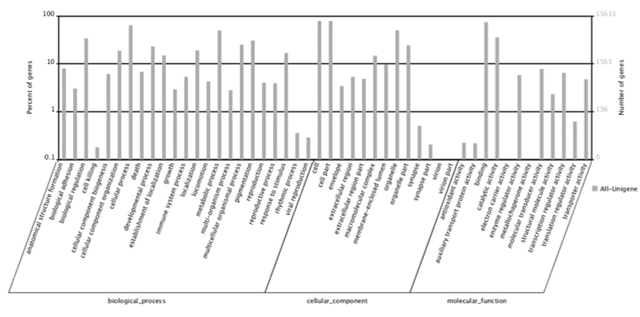
Gene Oncology (GO) and Clusters of Orthologous Groups (COG) ClassificationIdentified genes were categorized by function into orthologous groups by gene ontology (GO) assignments. Based on sequence homology, 15,633 sequences were categorized into 48 functional processes (Figure 2). The most common genes within the "biological process," "cellular component," and "molecular function" and "binding," respectively. Developmental process and response to stimulation are functions most pertinent to regeneration in axolotls; each consists of over 10% of biological processes (Figure 2). The remaining homology assignments concurred with experimental expectations that epigenetic and developmental functions are active during regeneration. Of the remaining functions noted, a minimal number of genes fell under the "virion," "virion parts," "electron carrier activity," and "metallochaperone activity" categories. While these functional processes were not expected, their roles should be studied further to discover whether they have prominent tasks in axolotl regeneration. Figure 2. Histogram Presentation of Gene Ontology Classifications. Genes were summarized into three primary categories: biological process, cellular component, and molecular function. The right y-axis indicates the number of genes and the left y-axis indicates the percentage of genes from the respective primary category.
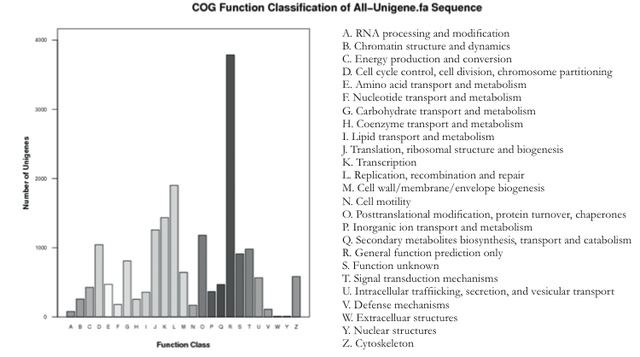
To test the accuracy of gene oncology assignments, assigned as well. The COG database provided a faster alternative for describing the functional characteristics non-redundant genes were categorized (Figure 3). Among the 25 COG categories, the clusters for "general function prediction only" represented the largest group (3785, 20.7%), followed by "replication, recombination and repair" (1901, 10.4%) and "transcription" (1436, 7.9%). In comparison to the COG analysis of the Taxus mairei genome, which showed "general function prediction only" as the largest group at 17.1%, it can be seen that the axolotl Taxus mairei (Hao et al., 2011). However, as "general function prediction only" is a poorly characterized group, it was excluded in the comparison. Otherwise, a notable difference was found in the "replication, recombination, and repair" synthesis, and an examination of cellular processes by gene emphasis of DNA regulation and cellular growth during axolotl regeneration. These annotations provide a valuable pathways during axolotl limb regeneration research. Figure 3. Histogram presentation of clusters of orthologous groups (COG) classification. Major functions are listed.
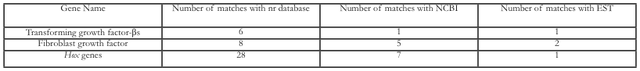
Table 2. Comparison of Selected Gene against nr, NCBI nucleotide, and EST databases. Three genes: transforming growth factor-s, hox genes were chosen for the comparison.
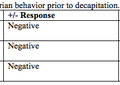
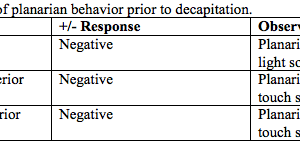
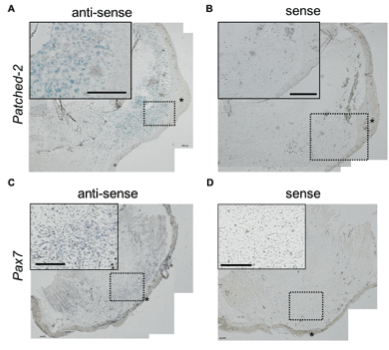
Detection of Sequences Related to Limb RegenerationFibroblast growth factors have been shown to contribute to vertebrate limb regeneration in axolotls (Poulin et al., 1993). For example, hoax-13, an autopod marker in many vertebrates, is expressed in the distal regions of the blastema (Satoh et al., 2011). Comparisons with the NCBI, EST, and nr databases yielded a multitude of sequences homologous to limb regeneration: transforming genes (Table 2). As tgfsequences (tgf-1, 2, 3, and 5) have been proven to be in the axolotl genome, the table captured a nearly complete portfolio of these genes and hox genes are linked to embryonic formation; their followed by redifferentiation into new limbs. Moreover, the high yield of high-quality sequences shows the ability of Illumina to facilitate research in axolotl genetics, whether previous genes. Figure 4. In situ hybridization analysis of patched-2 and pax7 expression in the regeneration blastemas. Patched-2 (2) or pax7 (C) signals (blue) using respective anti-sense probes are shown in the distal tips of the regenerating limbs. No signals were detected in the negative-control sections using respective sense probes of patched-2 (B) and pax7 (D) on serially sectioned slides. Scale bars indicate 200 μm.
Expression Patterns of Differentially-Expressed Genes in Limb RegenerationTechnical difficulties in the isolation of the regeneration blastema may produce inaccuracies, such as the accidental contamination of tissue samples by the underlying epidermis, which can lead to the higher expression of certain genes. Thus, in situ In situ hybridization showed that patched-2, one of the genes differentially expressed in blastema, is expressed in the cells located in blastema (Figure 4A ). Conversely, in situ hybridization with sense control probes gave no signals (Figure 4B). It can be concluded, then, that patched-2 expression is limited to within the blastema, and not in the surrounding epidermis (Figure 4A). Similarly, pax7 is also expressed only in the blastema (Figure 4C); pax7-positive cells also appeared in tail blastema and midbud-stage limb blastemas (Altschul et al., 1997). Therefore, the expression of regulatory genes involved blastema and not in surrounding non-blastemal tissues. These results demonstrate that the NGS libraries may identify potential markers for the blastemal progenitor cells that initiate limb regeneration. AcknowledgementsI would like to thank Professor HsuanShu Lee, M.D. for the opportunity to intern in his laboratory at National Taiwanese University Hospital during my half-year stay in Taiwan. I would also like to acknowledge Charlie Wu and the rest of the lab members for the oversight of my experiments in Professor Lee's lab. This study was supported by the National Science Council, Taipei, Taiwan. ReferencesAltschul, S., Madden, T., Schaffer, A., Zhang, J., Zhang, Z., & Miller, W., et al. (1997). Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Research, 25(17), 3389-3402. Bentley, D., Balasubramanian, S., Swerdlow, H., Smith, G., Milton, J., & Brown, C., et al. (2008). Accurate whole human genome sequencing using reversible terminator chemistry. Nature, 456(7218), 53-59. Campbell, L., Suarez-Castillo, E., Zuazaga, H. O., Knapp, D., Tanaka, E., during axolotl limb regeneration. Developmental Dynamics, 240(7), 18261840. Caron, H., Schaik, B. v., der Mee, M. v., Baas, F., Riggins, G., & Sluis, P. v., et al. (2001). The Human Transcriptome Map: Clustering of Highly Expressed Genes in Chromosomal Domains. Science, 291, 1289-1292 . Ferris, D. R., Satoh, A., Mandefro, B., Cummings, G. M., Gardiner, D. M., & Rugg, E. L. (2010). Ex Vivo Generation Of A Functional And Regenerative Wound Epithelium From Axolotl (Ambystoma Mexicanum) Skin. Development, Growth & Differentiation, 52(8), 715-724. Habermann, B., Bebin, A., Herklotz, S., Volkmer, M., Eckelt, K., & Pehlke, K., et al. (2004). An Ambystoma mexicanum EST sequencing project: analysis of 17,352 expressed sequence tags from embryonic and regenerating blastema cDNA libraries. Genome Biology, 5(R67), 1-2. Hao, D. C., Ge, G., Xiao, P., Zhang, Y., & Yang, L. (2011). The First Generation Sequencing. PLoS ONE, 6(6). Retrieved May 14, 2013, from http://www.plosone.org/article/info:doi/10.1371/jo Koniski, A., & Cohen, N. (1998). Axolotl (Ambystoma Mexicanum) Lymphocytes Produce And Are Growth-inhibited By Transforming Growth Factor-beta.Developmental & Comparative Immunology, 22(1), 91-102. Li, R., Zhu, H., Ruan, J., Qian, W., Fang, X., Shi, Z., et al. (2010). De novo assembly of human genomes with massively parallel short read sequencing. Genome Res, 20:265-272. Mchedlishvili, L., Epperlein, H. H., Telzerow, A., & Tanaka, E. M. (2007). A Clonal Analysis Of Neural Progenitors During Axolotl Spinal Cord Regeneration Reveals Evidence For Both Spatially Restricted And Multipotent Progenitors. Development, 134(11), 2083-2093. Monaghan, J., Epps, L., Putta, S., Page, R., Walker, J., & Beachy, C., et al. (2009). Microarray and cDNA sequence analysis of transcription during nerve-dependent limb regeneration. BMC Biology, 7(1), 179-198. Monaghan, J., Athippozhy, A., Seifert, A., Putta, S., Stromberg, A., regenerating limb of the Mexican axolotl. Biology Open, 1, 937-948. Muneoka, K., Han, M., & Gardiner, D. (2008). Regrowing Human Limbs. , 298, 56-63. Parchman, T. L., Geist, K. S., Grahnen, J. A., Benkman, C. W., & Buerkle, C. A. (2010). Transcriptome Sequencing In An Ecologically Important Tree Species: Assembly, Annotation, And Marker Discovery. BMC Genomics, 11(1), 180. Poulin, M., Patrie, K., Botelho, M. J., Tassava, R., & Chiu, I. (1993). during limb regeneration in newts (Notophthalmus viridescens). Development, 119(2), 353-361. Robertson, G., Chan, S. K., She, R., Varhol, R., Kamoh, B., & Prabhu, A. (2010). De Novo Assembly And Analysis Of RNA-seq Data. Nature Methods, 7(11), 909-912. Satoh, A., Graham, G., Bryant, S., & Gardiner, D. (2008). Neurotrophic Regulation Of Epidermal Dedifferentiation During Wound Healing And Limb Regeneration In The Axolotl (Ambystoma Mexicanum). Developmental Biology,319(2), 321-335. Satoh, A., makanae, A., Hirata, A., & Satou, Y. (2011). Blastema Induction In Aneurogenic State And Prrx-1 Regulation By MMPs And FGFs In Ambystoma Mexicanum Limb Regeneration. Developmental Biology, 355(2), 263-274. Smith JJ, Putta S, Zhu W, Pao GM, Verma IM, Hunter T, Bryant SV, Gardiner DM, Harkins TT, Voss SR., et al. (2009). Genic regions of a large salamander genome contain long introns and novel genes. BMC Genomics 10:19. Tamura, K., Yonei-Tamura, S., Yano, T., Yokoyama, H., & Ide, H. (2008). The Autopod: Its Formation During Limb Development. Development, Growth & Differentiation, 50, S177-S187. Yamada, K., et al. (2003). Empirical Analysis Of Transcriptional Activity In The Arabidopsis Genome. Science, 302(5646), 842-846. Suggested Reading from Inquiries Journal
Inquiries Journal provides undergraduate and graduate students around the world a platform for the wide dissemination of academic work over a range of core disciplines. Representing the work of students from hundreds of institutions around the globe, Inquiries Journal's large database of academic articles is completely free. Learn more | Blog | Submit Latest in Biology |