From Discussions VOL. 11 NO. 2Sodium Absorption in Summer and Winter Acclimated Freshwater Teleosts
By
Discussions 2015, Vol. 11 No. 2 | pg. 1/1
IN THIS ARTICLE
KEYWORDS
ABSTRACTTo maintain sodium (Na+) homeostasis in a hypotonic environment, freshwater teleosts must constantly absorb Na+ through their gills. Teleosts in temperate climates have the extra challenge of living in an environment in which ambient temperatures range from 0-30°C. It was hypothesized that 1. Na+ absorption through the gills occurs via a protein homologous to mammalian Na+/H+ exchanger (NHE), 2. Na+ is exchanged for NH4+—the nitrogenous waste product of the fish—rather than H+, and 3. gill proteins from 5°C winter-acclimated fish (WA) would be more active than those from 20°C Summer-acclimated fish (SA). To test this, the genome and transcriptome of SA fathead minnows (Pimephales promelas) were sequenced and assembled. RNA and DNA from SA were isolated using the Qiagen RNeasy Plus Mini-Kit, and the QIAamp DNA Mini-Kit, respectively. The tests found SAF expressed proteins homologous to four mammalian salt transport proteins: NHE, Na+/ Cl- co-transporter (NCC), acid-sensing ion channels, and NH3 transporters (Rh factors). Transcriptomes of SA and WA are now being sequenced the activity of Na + transport proteins in the gills measured. Preliminary data showed that Na+ flux into gill vesicles saturated with a K1/2 of 7.5 mM indicating a carrier-mediated process. Subsequent tests will measure the activities of NHE, NCC, and acid-sensing ion channels in SA and WA under various ionic conditions. IntroductionThere are many models for how freshwater (FW) teleosts maintain Na+ homeostasis in hypotonic environments. August Krogh was the first to propose a model for Na uptake. He discovered that Na+ and Cl- absorption were independent leading him to propose that Na+ was exchanged for NH4+ and Cl- for HCO3- (Krogh, 1939). Later tests by Avella and Bornancin (1989) showed that Na+ was more likely exchanged for protons than ammonia.
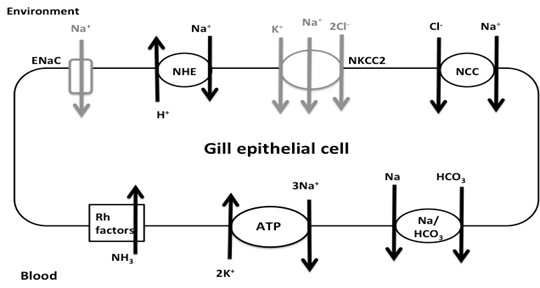
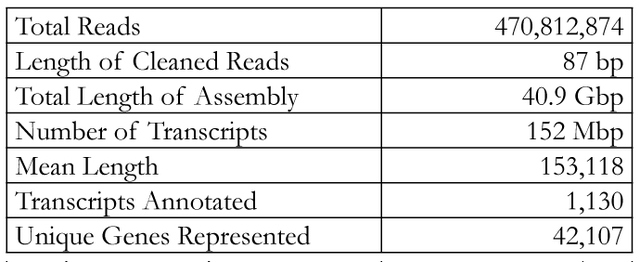
Figure 1. Na+ transporters of interest in warm acclimated fish. Transport proteins in black correspond to transcripts found in the gill transcriptome: NHE, NCC, Rh factors, NKA, and Na+/ HCO3- co-transporter. ENaC and NKCC2 are in grey since they were not found in the transcriptome. Recent research has shown there are three major models for Na+ uptake in FW fish: electrically coupled exchange using the epithelial Na+ channel (ENaC) linked to an H+-ATPase, uptake via Na+/H+ exchanger (NHE), and Na+ and Cl- co-transport (NCC) (Kumai & Perry, 2012). However, these models all have certain aspects that make them incomplete. For example, while the ENaC model works thermodynamically, there has been no evidence of ENaC, or its homologues, in sequenced FW fish genomes (Dymowska et. al, 2012). Genomic studies of the rainbow trout have shown the presence of NHE, but since it is driven by the concentration gradients of Na+ and H+, it should not be able to function in FW environments since extracellular Na+ is lower than intracellular Na+(Parks et al. 2008). One suggested way of overcoming the thermodynamics of NHE is through the use of ammonia transporters (Rh factors). Natawa and associates (2007) have proposed that Rh factor proteins transport NH3 out of the cell where it combines with H+ to form NH4+, causing an increase in local pH outside the cell making it favorable for Na+ transport via NHE. This model depends heavily on the creation of a microenvironment that is unstirred, but given the anatomy of the gills, and the high flow rate of water across the filaments, this unstirred layer is unlikely to exist. The last model, NCC, has only recently been identified in FW teleosts (Hiroi et al. 2008), but Na+ absorption via this transporter is also thermodynamically unfavorable given the extracellular and intracellular concentrations of Na+ and Cl- (Evans, 2011). It was proposed that Na+ absorption occurs by Krogh’s original model in which extracellular Na+ is exchanged for intracellular NH4+ and that NHE is the protein that facilitates this exchange. In addition Na+ absorption, FW teleosts in temperate climates must be able to maintain enzyme function from 0-30ºC. Since enzyme function falls with decreasing temperatures, such a large range poses a challenge for epithelial transport proteins maintaining Na+ homeostasis. Research by Packer & Garvin (1998) has shown that Na+/K+ ATPase (NKA) activity is higher in cold acclimated teleosts compared to warm acclimated teleosts when assayed at the same temperature. NKA moves Na+ from the intracellular milieu to the blood, thus generating the concentration gradient needed for Na+ entry. Possible causes for the change in activity include changes in protein expression, membrane lipids, or transport protein subunits. The Rosy Red fathead minnow (Pimephales promelas) was chosen for this study because of its use in ecotoxicology, commercial importance, and ability to tolerate environmental conditions under various temperatures, pH, and alkalinity. A member of the Cyprinidae family, the fathead minnow is broadly distributed across North America and is the model organism for aquatic toxicology studies and the Rosy Red strain is sold in pet stores as aquarium fish (Ankley & Villeneuve, 2006). It was hypothesized that 1. Na absorption through the gills occurs via a protein homologous to mammalian Na/H exchanger (NHE), 2. Na is exchanged for NH4, the nitrogenous waste product of the fish, rather than H+, and 3. Gill proteins from 5°C winter-acclimated fish will have higher activity than those from 20°C summer-acclimated fish (WA). This hypothesis was tested by sequencing the transcriptome of a SA Rosy Reds and measuring the activity of transporters potentially involved in Na+ re-absorption using fluorescence tagging. Materials and MethodsRNA PreparationRosy Red Fathead minnows were acclimated to 22ºC prior to tissue extraction. Fish were injected with heparin then anesthetized in MS-222 Tricane solution and dissected on ice. The opercula were removed and the ventral abdomen opened from anal fins to gills. Tissue and bone were removed to expose the ventricle. A 30G needle was inserted into the conus arteriosus, then pushed into aorta and tied in with 8-0 suture to prevent back flow, and the gills were flushed with 6mL of physiological saline and heparin. Gill filament RNA was extracted using the Qiagen Rneasy® Plus Mini Kit by manufacturer’s protocol (Cat. No. 74134, Qiagen, Hilden, Germany). Two samples were made, one from the right gill basket and the other from the left gill basket. RNA content and quality was determined using a Nanodrop Spectrophotometer, RNA samples were kept at -80ºC until sequenced. RNA Sequencing, assembly, and annotationRNA sequencing and library preparation was performed by the Case Western Reserve University Genome Sequencing Core using an Illumina HiSeq 2500 (Illumina, San Diego, CA, USA). Samples of 14.0 μg and 7.6 μg of RNA were used to construct two cDNA libraries using an Illumina TruSeq Stranded mRNA Sample Prep Kit (Illumina, San Diego, CA). Sequencing was done in two lanes, one for each sample, producing 101 bp paired reads for each sample. Raw reads were stored as FastQ files. George Washington University’s Colonial One High Performance Computing Initiative was used to assemble and annotate the transcriptome. For quality control, the first and final thirteen base pairs of each scan were removed using the Fasts Toolkit (Hansen, et al. 2010). Both samples were then combined and de novo assembly performed with The Broad Institute’s Trinity software package using default settings (Grabherr et al. and Haas et al.). The resulting transcripts were passed to the Broad Institute’s TransDecoder software and transcribed into protein sequences. The majority of functional gene annotation was done through the Broad Institute’s Trinotate Software package and protocol. Other software, including WEGO gene ontology and Kyoto Encyclopedia of Genes and Genomes (KEGG), was used to create a more complete annotation. All references to default parameters refer to the parameters given in the Trinotate protocol, or when none are specified the defaults of the software itself. Statistical analysis of the output files was done using scripts provided by the Trinity software package. Membrane Vesicle PreparationVesicle preparation was adapted from Flik and associates (1997). All vesicle preparation was performed at 4°C. After anesthetization in MS-222, the gills from two WA were homogenized in 1 mL of homogenizing solution which contained 50mM Sucrose, 10mM Tris/Hepes buffer 1:1, 1mM dithiothreitol, 1 mM ethylenediaminetetraacetic acid (EDTA). The resulting solution was centrifuged at 5,000xG for 15 minutes at 4°C. The supernatant was then spun at 50,000xG for 60 minutes. The pellet was resuspend pellet in 1mL sucrose solution which contained 300mM Sucrose, 10mM Tris/Hepes buffer 1:1, 1 mM EDTA and split into two 500uL samples. In one sample Na Green, a Na+-sensitive fluorescent dye, was added to make a 1μM final concentration and the other sample received the same volume of sucrose solution. Both were run through a 26.5G needle 20 times to allow the new solutions to be taken up by the vesicles. The solution was then spun at 50,000xG for 60 minutes to collect the vesicles. The pellets were finally suspended in 1.2 mL sucrose solution. Table 2. Summary of transcriptome analysis. Over 42,000 unique genes were found, and the high mean length indicates a transcriptome of high quality. FluorescenceEach sample was placed into a 3 mL cuvette with stirring. Over the course of 60 seconds, volumes of 150 mM NaCl were added corresponding to final Na+ concentrations of 2, 4, 6, 8, 16, and 32 mM and the change in fluorescence measured. Results & DiscussionThe results of sequencing the gill RNA of P. promelas using an Illumina HiSeq 2500 resulted in high quality sequencing and assembly, as determined by the large mean length of each transcript (Table 2).
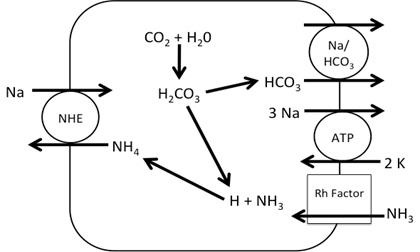
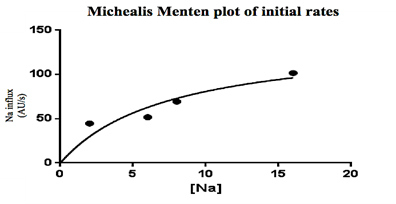
Figure 2. Working Model of Na+ absorption. NH3 is taken into the cell via Rh factor proteins where it combines with H+ from carbonic acid in the cell making NH4+. The NH4+ is used to in place of H+ in NHE allowing Na+ uptake into the cell. While the entire transcriptome was annotated, transporters related to Na+ uptake were detected: namely, transcripts corresponding to NHE, NCC, Na/HCO3 and Na/K ATPase (NKA). No transcripts related to ENaC or the Na/K/Cl cotransporter (NKCC) were detected. Initial query showed 14 different Rh factor proteins present in the transcriptome though due to false positives this number may be lower (Figure 1). From the transporters found in the transcriptome, a working model was developed for sodium entry. In this model, NH3 is taken into the cell via Rh factor proteins where it combines with H+ from carbonic acid in the cell making NH4+. The NH4+ is used in place of H+ in NHE thus allowing Na+ uptake into the cell. Na+ is then transported into the blood via NKA and the Enzyme activity of gill membrane vesicles containing the sodium-sensitive fluorescent dye, Sodium Green, was measured. Using the initial rates from each concentration, a Michealis-Menten plot was created where Vmax=141 and K1/2 = 7.5. These indicate that rises in fluorescence were due to carrier-mediated transport of Na+ into the vesicles (Figure 3).
Figure 3. Calculated rate of Na influx as a function of extra-vesicular Na (in mM) in warm acclimated fish. The Vmax=141 and the K1/2= 7.5. These data suggest the existence of Na+/H+ exchanger, Na+/HCO3-, Na+/Cl- co-transport, Na+/K+ ATPase, and NH3 transporters as these were identified in the gill transcriptomes but neither the ENaC nor NKCC2 were present in the transcriptomes. Regardless, Na+ influx kinetics remained consistent with NHE as a mechanism of entry. With these data, it is possible to construct a novel working model for how Na+ is absorbed through the gill epithelium. Future studies should focus on differential expression between SA and WA gill transcriptomes to identify differences at the mRNA level. These differences can then be linked to changes in function using fluorescent tagging. AcknowledgementsI would like to thank my Dr. Jeffrey Garvin, Dr. Randall Packer, Varsha Aravindabose, and Simon Wentworth for their contributions and support of this project. ReferencesGrabherr MG, Haas BJ, Yassour M, et al (2011) Full-length transcriptome assembly from RNA-Seq data without a reference genome. Nat Biotechnol 29: 644-652. doi: 10.1038/nbt.1883. Avella M, Bornancin M (1989) A new analysis of ammonia and sodium-transport through the gills of the fresh-water rainbow trout (Salmo Gairdneri). J Exp Biol 142:155-175. Dymowska AK, Hwang P, Goss GG (2012) Structure and function of ionocytes in the freshwater fish gill. Respiratory physiology & neurobiology 184: 282-292. doi: 10.1016/j.resp.2012.08.025 Evans DH (2011) Freshwater fish gill ion transport: August Krogh to morpholinos and microprobes. Acta physiologica (Oxford, England) 202:349-359. doi: 10.1111/j.1748-1716.2010.02186.x Flik, G., Kaneko, T., Greco, A. M., Li, J., & Fenwick, J. C. (1997). Sodium dependent ion transporters in trout gills. Fish Physiology and Biochemistry, 17(1), 385-396. doi:10.1023/A:1007768825043 Haas BJ, Zody MC (2010) Advancing RNA-Seq analysis. Nat Biotechnol 28: 421-423 doi: 10.1038/nbt0510-421 Hansen KD, Brenner SE, Dudoit S (2010) Biases in Illumina transcriptome sequencing caused by random hexamer priming. Nucleic Acids Res 28:e131. doi: 10.1093/nar/gkq224 Hiroi J, Yasumasu S, McCormick SD, Hwang P, Kaneko T (2008) Evidence for an apical Na-Cl cotransporter involved in ion uptake in a teleost fish. J Exp Biol 211:2584-2599. doi: 10.1242/jeb.018663 Krogh A, 1874-1949 (1939) Osmotic regulation in aquatic animals, England; United Kingdom Kumai Y, Perry SF (2012) Mechanisms and regulation of Na(+) uptake by freshwater fish. Respiratory physiology & neurobiology 184:249-256. doi: 10.1016/j.resp.2012.06.009 Nawata CM, Carrie C. Y. Hung, Tommy K. N. Tsui, Wilson JM, Wright PA, Wood CM (2007) Ammonia excretion in rainbow trout (Oncorhynchus mykiss): evidence for Rh glycoprotein and H+-ATPase involvement. Physiological Genomics 31:463-474. doi: 10.1152/physiolgenomics.00061.2007 Packer RK, Garvin JL (1998) Seasonal differences in activity of perch (Perca flavescens) gill Na+/K+ ATPase. Comparative Biochemistry and Physiology Part B: Biochemistry and Molecular Biology 120:777-783. Suggested Reading from Inquiries Journal
Inquiries Journal provides undergraduate and graduate students around the world a platform for the wide dissemination of academic work over a range of core disciplines. Representing the work of students from hundreds of institutions around the globe, Inquiries Journal's large database of academic articles is completely free. Learn more | Blog | Submit Latest in Biology |