Mycobacteriophage Meru: Isolation and Characterization of a Novel Mycobacteriophage
By
2013, Vol. 5 No. 10 | pg. 1/3 | »
IN THIS ARTICLE
KEYWORDS
Mycobacteriophage are the most prevalent type of microorganism present in the biological universe. In fact, since the first mycobacteriophage was isolated in 1947, well over 2000 new viruses have been identified (Mycobacteriophage Database). The vastness in number lends itself to widespread genetic diversity and variation, making phage research one of the most compelling modern-day fields of research in microbiology. Specifically, the recognition of the immense numbers of bacteriophages in the planet has contributed to a renewed interest in understanding their morphological and genetic diversity, and explaining the evolutionary mechanisms responsible for such variation (Hatful, 2008). With this notion in mind, the Howard Hughes Medical Institute’s Science Education Alliance (SEA) recently began the National Genomics Research Initiative, with the aim of significantly expanding the available data on mycobacteriophage. As part of this initiative, students at Virginia Commonwealth University collected dirt samples from various locations, and individually commenced the process of isolating and characterizing a mycobacteriophage for eventual genome sequencing. See further details about Phage Meru at the Mycobacteriophage Database: http://phagesdb.org/phages/Meru/ Specific procedures involved in the characterization of a mycobacteriophage are exceptionally useful in comparing behavior between different phages. A chief example is cross- immunity testing, in which mycobacteriophage are spotted onto bacterial lysogens, to test for super-infection immunity. This methodology is particularly useful to identify the specific life cycle of the mycobacteriophage from two distinct types that have been identified through prior phage research: lytic and lysogenic. According to the Mycobacteriophage Database, a phage that follows a lysogenic cycle injects its genetic material into the bacterial genome, but does not immediately lyse the bacterial cell. Typically, the phage genome becomes fully integrated with the bacterial genome, and future replication of the bacterial chromosome involves the combined genetic mass. On the other hand, the lytic cycle of a phage is characterized by the injection, replication, and expression of the phage genome, in addition to the thorough lysis of the bacterial cell as the phage exit to find a new host cell and begin a new round of infection. (Mycobacteriophage Database Glossary). Specifically, when testing for phage immunity, we can identify a certain mycobacteriophage as following a temperate cycle if the putative lysogen host bacterial cells show little or no cell lysis after being infected with the virus. This indicates that the bacterial cells were host to phages that could integrate themselves into the bacterial DNA and confer immunity to subsequent infection (Budzik, 2000). Genomic sequencing of mycobacteriophage such as L5 have indicated that a certain amino acid product of the L5 gene 71, also known as the repressor gene, is responsible for conferring immunity to L5 super-infection (Donnelly,1993). Accordingly, further genomic sequences of other temperate phages would reiterate the role of the repressor in stabilizing the lysogenic condition, as well as suggest evolutionary connections and patterns between temperate phages. Host range testing is another useful procedure that can serve as the basis for comparisons involving plating efficiency of different mycobacteriophage on different bacterial strains. Early phage research was hindered by the fact that the wide host range of the first phages limited their value in taxonomic studies (McNerney, 1999). However, the increase in the sheer number of identified mycobacteriophage indicates that host range analysis is more beneficial today, as relations between phages have appeared more frequently. Consequently, the main value of the procedure centers on the ability to form preliminary groups of related phages based on similar infection efficiencies on the same bacterial strain. In turn, further study can aim to identify additional similarities between these grouped viruses, including applications to mycobacterial genetic manipulation. The purpose of the current investigation is to present findings related to the isolation and initial characterization of a novel mycobacteriophage, Meru. Data pertaining to morphology, gel electrophoresis, cross-immunity testing with other mycobacteriophage in the VCU Phage Lab course, and host-range testing with two different strains of M. smegmatis are presented in an orderly fashion. Results indicate that Meru is a potential Cluster A temperate phage, with extremely turbid plaques that demonstrate mixed morphology. Immunity testing revealed that the Meru lysogen was not immune to any of the other mycobacteriophage from the course. Though the ability to test infection efficiency on a more diverse array of bacterial populations, including M. tuberculosis, was stymied by the necessary safety precautions of a classroom laboratory environment, host-range testing yielded curious results. Although Meru was isolated on lawns of the mc2155 strain of M. smegmatis, the virus also infected the ATCC type M. smegmatis strain with a uniquely heightened plating efficiency. MethodologyThe process of collecting samples of dirt and subsequently using specific methods to isolate a pure mycobacteriophage from the mixture spanned a period of several weeks. Dirt specimens were collected from a wooded area in Mineral, Virginia (Figure 1), the same region that was the epicenter of the earthquake that hit Virginia on August 23, 2011. 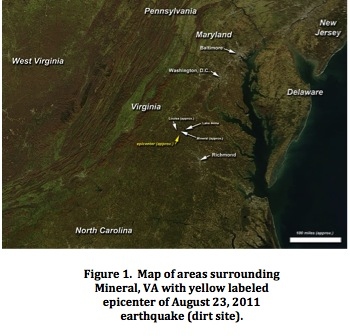 Next, an enrichment culture was created in order to provide an environment that favored the replication of bacterial phages. The culture included 7H9/glycerol broth, 1% albumin, 0.4% dextrose, 29 mM NaCl, 1 mM CaCl2 and an aliquot of 5 ml of stationary phage M. smegmatis culture. The flask was incubated, shaking at 220 rpm for 24 hours. Then, a phage filtrate was prepared by filter sterilizing the supernatant using a 0.22 mm filter. The filtrate was diluted by a serial dilution from 100-10-4. Directly after the dilution was performed, previously prepared M. smegmatis culture tubes were infected with the serial dilution samples, vortexed well, and then set to rest for about twenty minutes. The infected tubes were then plated onto standard Luria Agar plates with top agar solution (7H9 broth base, 0.4% agar, 1 mM CaCl2). The plates were incubated for 48 hours at 370C. Four rounds of purification, which involved picking plaques with sterile tips, performing a ten-fold serial dilution from 100-10-4, and infecting and re-plating M. smegmatis culture tubes with top agar, were then conducted in order to determine if a pure phage population was isolated. Two stick streak tests, in which sterile tips were used to streak a single, well-isolated plaque across three labeled sections of a plate, were also utilized to confirm the presence of an isolated population. Based on careful measurement of plaque and plate diameter on a single plate, empirical testing was done to predict the concentration of phage to plate to produce a ‘webbed plate’ with a lawn that is almost completely lysed. Consecutive rounds of serial dilution were performed with the purpose of achieving web patterns of lysis on bacterial lawns grown in top agar. Once web patterns were found, each plate was flooded with phage buffer (PB: 10 mM Tris, pH 7.5, 10 mM MgSO4, 68 mM NaCl, 1 mM CaCl2), and the contents were subsequently gathered for lysate collection via filter-sterilization. From the resulting high titer lysate, a fresh round of serial dilution was performed and the titer was calculated. Phage genomic DNA was then isolated and purified through a multi-step process. First, 10 mL of the filter-sterilized phage lysate were transferred into an Oak Ridge tube. Then, 40 microliters of nuclease mix (0.25 mg/ml RnaseA, 150 mM NaCl, 50% glycerol) were added to the tube in order to degrade the bacterial host DNA, followed by a thirty minute period of incubation at 370C and an hour of exposure to room temperature. Subsequently, phage genomic particles were precipitated from the nuclease-containing lysate by adding 4.0 mL of phage precipitant solution (30% PEG 8000, 3.3 M NaCl) and conducting thirty minutes of incubation and high-speed centrifugation. After decanting the supernatant and re-suspending the resultant pellet, DNA was purified using the materials and protocol recommendations provided in the Promega Wizard DNA purification kit (Madison, WI). Directly after purification procedures were completed, the concentration of the collected DNA was determined using a Nanodrop spectrophotometer. To aid in the identification of a unique mycobacteriophage to sequence and genetically analyze, restriction digest procedures were initiated by using restriction enzymes (ClaI and HaeIII) to cut the phage genomic DNA sample. Using gel electrophoresis, in which agarose gels of 1% and 1.5% concentrations volts were used to isolate and enable visualization of the bands with UV light, restriction patterns of the mycobacteriophage can be compared with those of known viruses, so as to determine the relative novelty of the current phage under study. Finally, about 10 microliters of phage lysate were mounted onto electron microscope grids coated with formvar, and stained with 1% uranyl acetate, in order to prepare for the observation of the mycobacteriophage using electron microscopy. The specific microscope utilized was a Jeol JEM-1230 TEM with a Gatan Ultra Scan 4000 camera using 120 kV for a brighter image. In order to assess the integration of phage DNA into bacterial genomes, a super- plaque was constructed by placing 10 microliters of the high titer lysate on a lawn of M. smegmatis. Lysogen streaking was then performed by gently scraping the developed super-plaque and streaking for bacterial survivors on a separate plate. Growth on this second plate was used to examine the putative lysogen’s immunity to super-infection by the mycobacteriophage. A liquid lysogen culture was then created by using a sterile tip to scrape part of the lysogen streak and inoculating a liquid 7H9 media culture. Lysogen growth in this media was plated using top agar and then served as the basis for spot-testing phages for immunity testing and cross-immunity analysis, two methods used to compare different mycobacteriophage--both lytic and lysogenic---and their infection efficiencies relative to each other’s lysogens. Finally, host range testing was also performed in order to assess and compare the infection efficiencies of the mycobacteriophage Meru on lawns of two different bacterial strains of M. smegmatis. To achieve this purpose, the phage high titer lysate underwent a full round of serial dilution and was spotted onto two plates pre-loaded with top agar and either the mc2155 strain or the ATCC type strain.Continued on Next Page » Suggested Reading from Inquiries Journal
Inquiries Journal provides undergraduate and graduate students around the world a platform for the wide dissemination of academic work over a range of core disciplines. Representing the work of students from hundreds of institutions around the globe, Inquiries Journal's large database of academic articles is completely free. Learn more | Blog | Submit Latest in Biology |