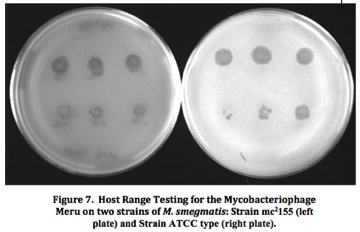
Mycobacteriophage Meru: Isolation and Characterization of a Novel MycobacteriophageResultsFrom the initial plating of M. smegmatis enrichment cultures, plaques were identified. The plate contained plaques of both small and large sizes, ranging from a diameter of about 0.5 mm to about 3 mm. All plaques appeared significantly light and turbid, and contained smooth edges. Four consecutive rounds of purification, which included separate purification dilutions for a phage with a diameter of 0.5 mm and a phage with a diameter of 3mm, yielded a consistent picture of plaque morphology (Figure 2). Each plate that resulted from these processes showed mixed plaque sizes within a similar range (0.5 to 3.0 mm). 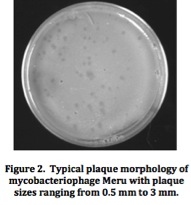 To confirm this consistency, two stick streak tests were utilized, which also yielded the same diversity in plaque sizes as well as similarities in the other characteristics of plaque morphology mentioned previously. As a result, we concluded that the mycobacteriophage isolated was of a pure population species because of persistent patterns in size, shape, turbidity, and general structure. Empirical testing for web patterns yielded plaque morphologies that were consistent with previous purification results (mixed plaque sizes ranging from less than 1 mm to about 3 mm; high turbidity) (Figure 3).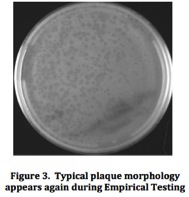 The titer calculation that was made upon serial dilution and plating of M. smegmatis infected by filter-sterilized high titer phage lysate yielded a result of 4.7 x 108 pfu/mL. Subsequent to phage DNA isolation and purification, the concentration of phage DNA was found to be 86.6 ng/µL. Because of the relatively small concentration of DNA retrieved from DNA isolation and purification methods, gel electrophoresis was only conducted with two restriction enzymes (ClaI and HaeIII). Results of the procedure (Figure 4) indicated that ClaI was not able to cut the DNA, while HaeIII was able to digest the genomic material. 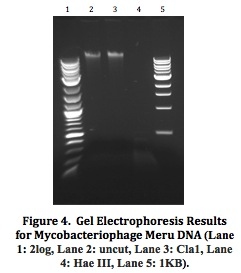 Cross-immunity testing revealed that Meru was not immune to any of the other phages in the Fall 2011 Virginia Commonwealth University Phage Discovery course. However, there appears to be doughnut shaped accumulation in some of the spots, where lysis seems to be disrupted (Figure 5). The sequenced phages Batiatus (Cluster F) and Twister (Cluster A1) were able to lyse the Meru lysogen, indicating that Meru may be unrelated to phage from these commonly isolated clusters. Immunity testing revealed that the mycobacteriophage Tibbie, a potentially lytic phage from the course, significantly lysed the mycobacteriophage Meru (Figure 6). Finally, it is interesting to note the conversion of the Meru lysogen lawn to a more lytic condition, as observed by formation of plaques within the bacterial lawn. 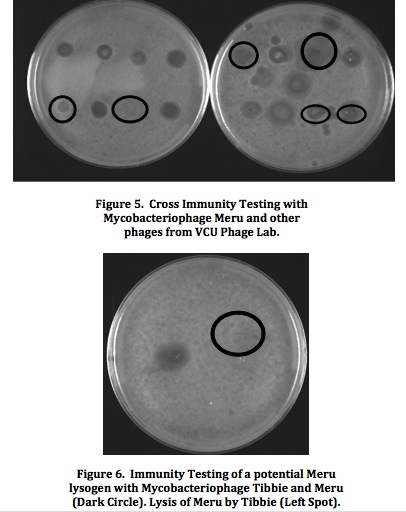 Host range testing yielded similar plating efficiencies for the mycobacteriophage Meru on both strains of M. smegmatis (Figure 7). All plaques that appeared were turbid, especially those plated from the latter portion of the serial dilution. In terms of texture and size, the plaques were mostly smooth-edged and very consistent in diameter. However, it appears that Meru had a greater infection efficiency on ATCC type strain for the last few spottings, as evidenced by a slightly greater degree of lysis in those areas of the plate.
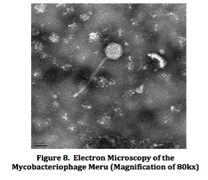
Lastly, electron microscopy provided a clear picture of the physical structure of Meru (Figure 8). Using a scale marker of 50 nm, the tail length of the phage was calculated to be about 204 nanometers, while the head diameter was found to be about 69 nanometers.Continued on Next Page » Suggested Reading from Inquiries Journal
Inquiries Journal provides undergraduate and graduate students around the world a platform for the wide dissemination of academic work over a range of core disciplines. Representing the work of students from hundreds of institutions around the globe, Inquiries Journal's large database of academic articles is completely free. Learn more | Blog | Submit Latest in Biology |