Studying Ancient DNA to Understand Contemporary Disease
By
2018, Vol. 10 No. 12 | pg. 1/1
IN THIS ARTICLE
KEYWORDS
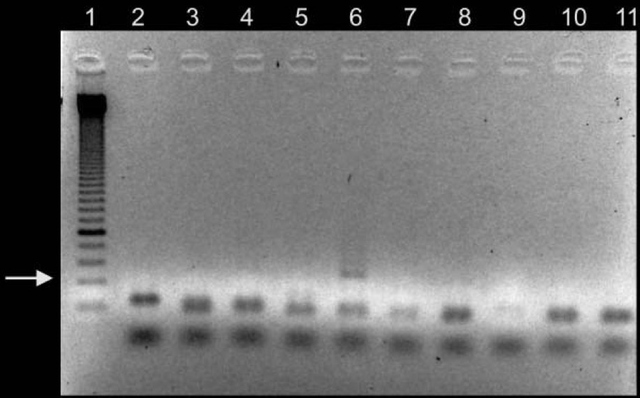
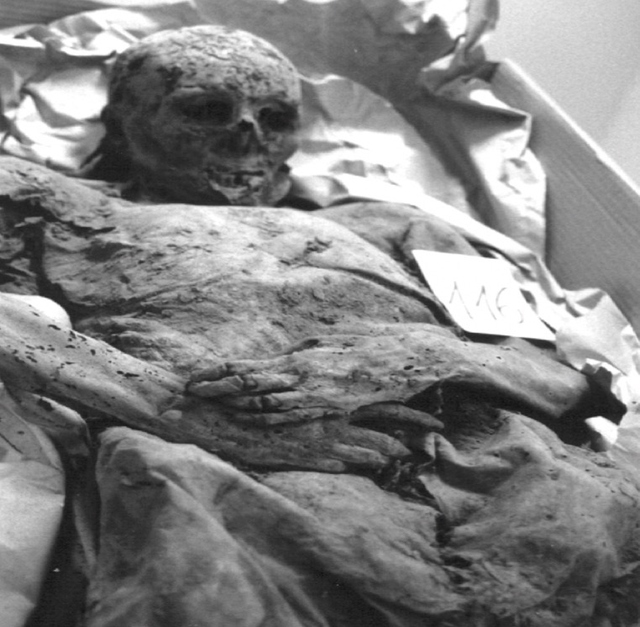
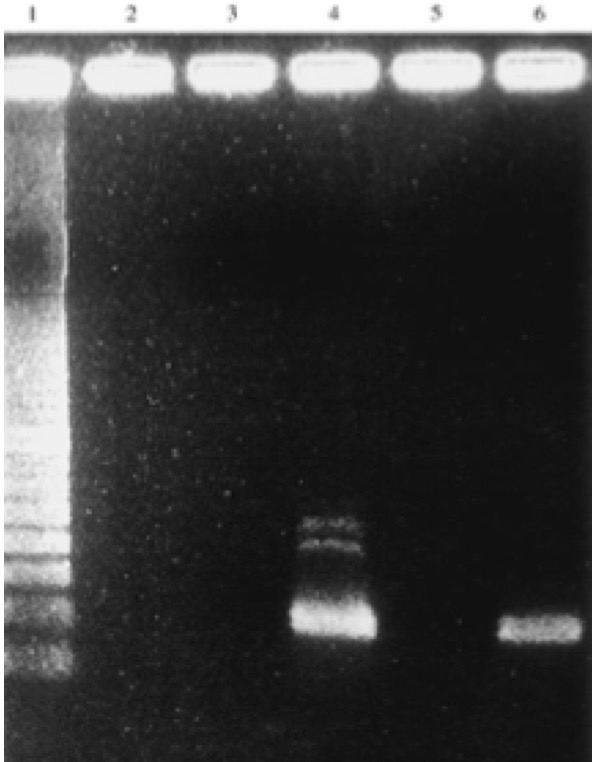
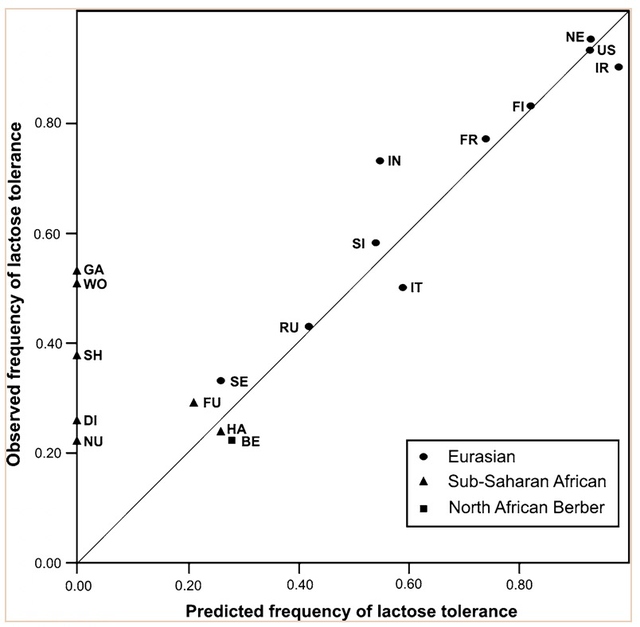
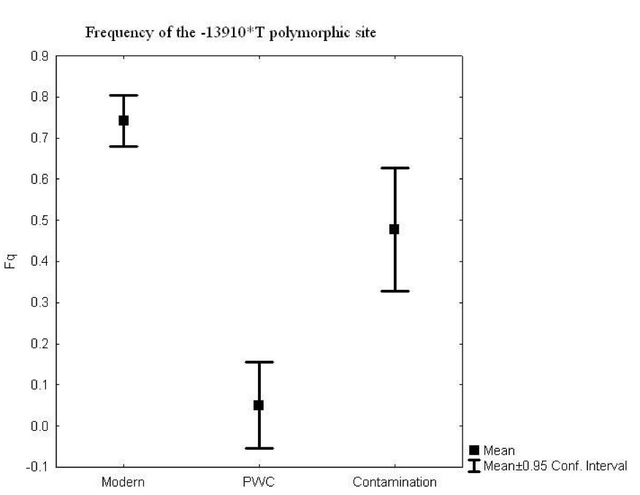
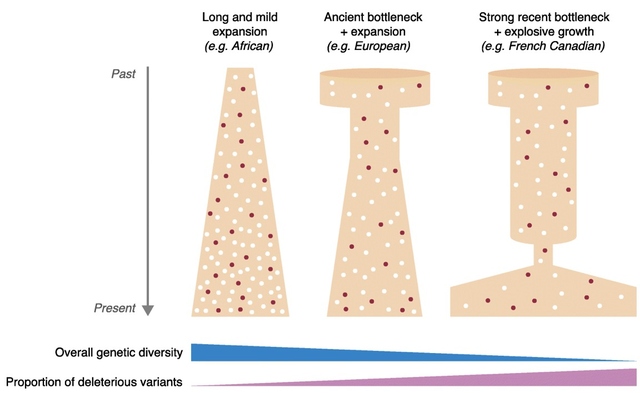
The study of DNA and genetics has always been a large mystery to many scientists. The current Ancient DNA (aDNA) research on human history is more complex than what can be inferred from modern DNA research. Scientists and researchers are constantly using modern day populations, and modern DNA to make inferences about past populations (Haber et al., 2016). With the new technologies available in ancient DNA, the study of past diseases and populations is more easily conducted with little to no contamination. Studying aDNA does not only tell us about current and past disease, but can also shed light on the theories of human origins (Haber et al., 2016). For instance, there is lots research done surrounding the out of Africa hypothesis and more recently the leaky replacement model (Haber et al., 2016). These are some of the new research ideas that are being introduced from aDNA studies. Ancient DNA research is changing the views of human origins by displaying one that is more complex through looking for evidence from aDNA and interpreting with modern day genetics (Haber et al., 2016). In addition to using aDNA for discoveries about human origins, it is also used for inferences on the spread of new diseases based on ancient evidence. Ancient DNA can be used to study ancient diseases such as tuberculosis and malaria. Microbial DNA is the main component that will be interpreted in terms of aDNA. Due to external factors such as contamination, the ability to distinguish between ancient and modern DNA makes it difficult to determine disease origins and susceptibility (Haber et al., 2016). This paper will look at microbial DNA and how it can be used to make inferences on past and present diseases, ancient DNA and malaria, tuberculosis and lactose gene mutation. Issues with using aDNA will be addressed because they play a fundamental role in understanding aDNA extraction and analysis methods. These are some of the many components in aDNA that will be looked at because it sheds light on how it plays a role on disease past and present. Disease patterns using aDNA are fundamental and this can allow for the understanding of origins and further disease patterns. Ancient Egypt and MalariaPlasmodium falciparum (P.falciparium) is also known as malaria and is caused by the single cell parasite known as plasmodium. In modern times, malaria is endemic in the African continent due to climate and environmental changes. With evidence of malaria presence in Ancient Egypt and Greece it origins can be traced, thus this can provide an insight on modern patterns in endemic countries (Nerlich et al., 2008). Plasmodium falciparum is an infection caused by mosquitoes, which causes clinical infection (Nerlich et al., 2008). The others include, plasmodium malariae, plasmodium Vivax and plasmodium ovale (Nerlich et al., 2008). These types of malaria infections lead to different symptoms thus rendering them different from the well-known malaria.The study done by Nerlich et al., (2008) states that the identification of aDNA for P.falciparium can be found in the tissue of an ancient mummy dating back to approximately 4000 years ago. Bone tissue samples were collected from 91 mummies and skeletons from Ancient Egypt. The samples were analyzed using PCR for direct sequencing of P.falciparium. The results showed that 2 of the 91 individuals had the fragment of P.falciparium, which are 134 base pairs (bp) (Figure1). This fragment has a 99% accordance ultimately allowing for the identification of this by using aDNA (Nerlich et al., 2008). In addition, the samples from the individuals showed that they originated in different tomb areas in different kingdoms ranging from the Pre-dynastic to Early Dynastic or Middle kingdom (Nerlich et al., 2008). The result of this study is a representation of how aDNA can be used to trace disease origins and gain in depth information on a disease (Nerlich et al., 2008). This study also outlines the difference of using immunological tests versus aDNA. This is significant because a fault of previous immunological tests have mislabelled chronic anemia has positive for P.falciparium thus rendering aDNA testing to be more superior. Figure 1: 134bp fragment from ancient DNA of Plasmodium falciparum extracted from ancient Egyptian mummies: (Nerlich et al., 2008: Figure 1 pg. 318) Moreover, building from the notion of the presence of P.falciparium in Ancient Egypt Lalremruata et al., (2013) conducted a study that looked at the co-infection of TB and malaria. Ancient DNA in this case can be used to identify multiple cases of diseases, which can provide a greater insight into disease progression and origins. Lalremruata et al., (2013) conducted a study that looked at ancient DNA and P.falciparium in mummies from 1500-500 BC which included 18th Dynasty. The 196 bp AMA1, MSP1 and MTB complex were looked for 6 mummies for the identification of P.falciparium (Lalremruata et al., 2013). These were used to determine the presence of TB and P.falciparium. Out of the 6 mummies that were analyzed, two had cases of single malaria infections and the remaining four had representations of malaria and TB confection (Lalremruata et al., 2013). Lalremruata et al., (2013) argue that the Fayum was an area that was more susceptible to malaria than it is today. Fayum has undergone drastic changes throughout antiquity from how its was described by Herodotus in Ancient Greek, to the Ptolemies and ancient Rome. Studies have showed that cultivation of land by humans lead to swamps that are ideal breeding grounds for Anopheles mosquitos the vector of P.falciparium (Lalremruata et al., 2013). It can be inferred that ancient populations (1500-500 BC) of Fayum might have been at an increased risk of malaria due to the reclamation of land that was caused by cultivation (Lalremruata et al., 2013). Studies using molecular identification cannot only give us information about past incidence of disease but also the environmental influences. Ancient DNA and TuberculosisAncient DNA has allowed information to be obtained in real-time rather than using molecular calculations that rely on a molecular clock that uses mutation rates (Donoghue et al., 2004). In addition, as seen with malaria, aDNA can provide information about early agricultural practices and health status in relation to disease and diet (Donoghue et al., 2004). More importantly, when using aDNA to study contemporary diseases, contamination is an important concern that needs to be addressed in order to produce accurate results. The first finding of tuberculosis was in 1993 using aDNA in a human microbial pathogen (Donoghue et al., 2004). In the first report of aDNA of tuberculosis 11 specimens were analyzed and four dated to age from 300-1400 years ago tested positive for M.tuberculosis (Donoghue et al., 2004) .The target sites for these specimens were IS6110 for tuberculosis DNA. In order to identify tuberculosis in ancient specimens bimolecular evidence from mycolic acids and DNA were used (Donoghue et al., 2004). The reason that this is used to identify tuberculosis DNA is because point mutations of the DNA are rare and thus a reliable source. In addition, independent variation can be used to detect different target sequences on the tuberculosis genome. For example, a study done by Fletcher et al., (2003) 168 individuals from the 18th century Hungry were analyzed due to their preservation in mummy form. DNA extraction was done using IS6110 bp analysis and the presence of the MTB complex. Out of the 168 individuals, 93 had the target sequences for M. tuberculosis (Figure 2) (Fletcher et al., 2003). In addition 27 individuals were radiographed and 14/27 had potential lesions and 11/14 that had lesions has chest examinations were positive for MTB. The two methods of DNA extraction and radiographic examination work together to aid in the identification of tuberculosis. The DNA found in the was well preserved and can highlight the molecular epidemiology of infection on this community which can allow for comparison with MTB strains that exist in the current time (Fletcher et al., 2003) Figure 2: Mummified body from Vac Hungry. DNA from the chest detected M. tuberculosis was positive. (Originally from Fletcher et al., 2003): (Donoghue et al., 2004: Figure 4) In addition to looking at the genomic markers to identify tuberculosis, osteological evidence can provide valuable insight to the long-term trends of the disease. In some cases tuberculosis can cause manifestation on bone, which is seen in the spine known as Pott’s Disease (Gernaey et al., 2001). Gernaey et al., (2001) conducted a study on medieval individuals where mycolic acids were used as the biomarkers for the identification of tuberculosis as opposed to the mycobacterium tuberculosis complex and the IS6110 fragments. Their reasoning is that the target is not as specific and not always present in modern strains (Gernaey et al., 2001). Mycolic acids are long chain lipids, and in the case of tuberculosis are 3 hydroxy fatty acids that are substituted into the 2-position with a long aliphatic chain (Gernaey et al., 2001). Mycolic acid along with DNA evidence can effectively be used to confirm the osteological evidence of TB in bone (Gernaey et al., 2001). Also, mycolic acids are important because it allows for the comparison with ancient DNA and younger specimens because it can screen biomarkers in individuals younger than 250 years. In the study conducted by Gernaey et al., (2001) ribs were used for the identification of tuberculosis because they are most preserved as compared to other parts of the skeleton. Samples were taken from 30 skeletons dating back to approximately1000 years old (Medival) and are from Addingham, West Yorkshire. To determine the presence of tuberculosis, three tests were conducted that used IS6110 and mycobacterium tuberculosis complex mycolic acids. Three of the skeletons were used, specimen A134 was a male that had Pott’s disease and new bone formation on the ribs Two mid shaft fragments were taken and ground to powder for DNA extraction. Specimen A223 had no lesions of TB in the spinious process but had lesions on the ribs that were possibly caused by pulmonary TB. Two mid shaft fragments were also selected and ground into powder. The third specimen A162 had no lesions of TB. This specimen was used as the control comparison (Gernaey et al., 2001). Mycolic acid extraction, chromatography separation and DNA analysis where the methods that were used to look at the osteological evidence of TB. The results of the study showed only one of the specimens, A134, had positive identification of IS6110 primers (Figure 3). The other two did not produce any bands associated with the target mycobacterium complex. The other two specimens showed evidence of mycolic acids. This study provides evidence that there are other methods using osteology that can provide information on TB. For instance this study did not look at straight radiological evidence but DNA and mycolic acids, which is a reliable form of ancient TB diagnosis than 1S6110 (Gernaey et al., 2001). What can also be inferred is that mycolic acids can survive for more than 1000 years as seen in the medieval specimens (Gernaey et al., 2001). This study provides valuable evidence that looks at TB through a paleoepidemiological perspective because it provides evidence of the presence of TB and the ability to trace it back to antiquity. Figure 3: Amplification of 181 bp product of the target IS6110. Positive result only from one rib sample from one individual (Gernaey et al., 2001: Figure 1) Potts disease- Physical Representation of TuberculosisWhen analyzing osteological evidence for tuberculosis, Pott’s disease is most looked at because it has the most manifestation on bone. Manifestation on a bone is indicative of the presence of disease. Crubézy et al., (1998) conducted a study that attempted to extract and recover DNA from a 5400-year-old skeleton from Pre-dynastic Egypt. This individual suffered from spinal deformity, which is secondary tuberculosis identification on the vertebrae. Parts of the skeleton that were used for DNA analysis include collapsed vertebral bodies from the eighth to tenth thoracic vertebrae, and a proximal eighth left rib that have periosteal new bone formation (Crubézy et al., 1998). For the DNA extraction/amplification of the IS6110, insertion element was conducted, which is the isolation of the insertion sequence IS in this cause 6110 (Crubézy et al., 1998). In addition, mycobacterium DNA variation phylogenetic tree was constructed to determine the ancestral sequence and mutational events. The results of the study showed the morphology of the vertebral lesions are similar to modern day skeletons that have spinal tuberculosis. The pre-dynastic specimens had fusion of vertebrae, remodelling of the inferior articular surfaces of the apophyses indicate tuberculosis and long-term disease presence. The specimens also exhibited periosteal new bone formation, which indicates terminal infection, which is usually a response to injury or stimuli. This is indicative of disease because it shows evidence of trauma, which could be from Potts disease. Crubézy et al.,(1998) argue that there is unusual diversity of their DNA sequence, which could be due to the large amount of damage in the ancient DNA and or the origins of the disease. For instance, the authors speculate that the agent of human tuberculosis arose from the cattle pathogen M.bovis. This could have resulted from damage to the DNA causing the human form of the disease. Also, the authors speculate that at this time there was long evolution of the disease due to the healed cases of bone, which could have resulted from an immunised population (Crubézy et al.,1998). Based on the periosteal new bone formation, there is evidence of healing due to previous exposure to TB. This would mean that the population would have has a chance to be naturally immune because of their encounter with the disease as seen on the specimens. Crubézy et al., (1998) cannot agree if the lesions on the individual were from M.tuberculosis or from M.bovis or an ancient mycobacterium that is similar to the two pathogens. The authors conclude that tuberculosis in humans cannot be less than 15000 years old. Ancient DNA and Lactase Gene MutationIn contrast to studying the genetic components of contemporary disease in ancient context, it is also important to understand gene mutations and how it has played a role in modern disease and dietary practices. This section will look at the human genome to study gene mutations such as lactose intolerance. More specifically, analyzing the lactase gene will allow a more thorough understanding of the origins of lactose intolerance. In a study done by Myles et al., (2005) pastoralism was analysed as a potential cause of the spread of lactase gene mutation in the Neolithic. This was explored through three North African Berber populations. The underlying notion is that the expansion of pastoralism from the Middle East into North Africa would have been the cause of the spread of lactose intolerance (Myles et al., 2005). The study looked at a 105 Berber samples that were from three different groups in Morocco and Algeria. To determine the lactase tolerance in the population, haplotype frequencies from 11 polymorphic sites were extracted and compared to other populations worldwide (Myles et al., 2005). The two polymorphic sites that were most causal for lactose tolerance were the -13910T allele and the -22018A allele (Myles et al., 2005). The findings show that individuals that carry the -13910T always carry the -22018A but not vice versa. From this knowledge individuals who carry the -22018A allele are lactose intolerant but allele does not affect the lactase promoter. The likely causal allele is -13910T, which was the main focus for the analysis of lactose tolerance. The allele -13910T is important when looking at the origins of the dairy culture (Myles et al., 2005). In previous studies it has been used to predict the frequency of lactose tolerance in Northern Europeans but not in Sub-Saharan countries due to the migration of populations. Myles et al., (2005) propose that the presence of the -13910T allele is related to lactose tolerance in Eurasian and Berber populations but absent from sub-Saharan African populations supports the hypothesis that there is shared dairying origins (Myles et al., 2005). Figure 4: Correlation between the frequency of lactose tolerance measured by a lactose test and lactose tolerance predicted by the Hardy-Weinberg equilibrium. The graph has abbreviations, which were explained as follows; NE (Northern Europe), US (United States), IR (Ireland), FI (Finland), FR (France), IN (Northern India), SI (Sindi, Pakistan) and many more global populations (Myles et al., 2005: Figure 2). The above graph is a representation of the frequency of lactose tolerance represented by the frequency of the -13910T allele in Berber and Eurasian populations. The diagonal line is a representation of a perfect correlation, which is not expected by the authors because the data was collected from different sources of genotypic data. This graph is important when looking at the origins of the gene mutation of lactase because it allows inferences on the origins of the mutation. This is done looking at small population groups and their genetic history. The lactase mutation is important in understanding human evolution because it provides insight in to the behavioural patterns of past populations. The analysis of the -13910T allele with its presence or lack of on the A haplotype can provide evidence for the origins of a population’s dairying pattern. The A haplogroup was observed on populations that were Europeans and was observed in eight of the studied individuals. The data can infer that -13910T is a young mutation and was not widely geographically spread (Myles et al., 2005). The direct causal influence of lactose tolerance is not definite but is believed to be associated with the domestication of ovicaprids (sheep and goats) (Myles et al., 2005). It is also believed that the change in pastoralism and dairying patterns was abrupt in North Africa. Myles et al., (2005) suggest that the proto-speaking Berber ovicaprid pastoralists were the cause of the introduction of the -13910T allele, ultimately causing lactose tolerance. This suggests that there was a genetic input from the Eurasian populations causing lacto. Looking at the lactase gene mutation can shed light on why there are so many individuals today who are both lactose tolerant or intolerant and how it potentially occurred. Lactase and Prehistoric PopulationsMoreover, while its important to look at modern populations when analyzing lactase gene mutation, prehistoric populations can provide insight on the interactions of genes and culture. Malmström et al., (2010) looked a Neolithic hunter-gather population in Northern Europe and the frequency of lactose intolerance as opposed to the ability to digest milk sugar lactose after childhood. The population is known as the Pitted-Ware culture (PWC) and were thought to be present in Scandinavia 5400-4300 years before present time (Malmström et al., 2010). The authors found in this study that a genetic component affecting the ability to digest milk as an adult would have resulted from the replacement of the population with an agricultural based population. This would have resulted from changing to a population that used cultivation and farming as opposed to hunter-gather group. This study looked at samples of teeth and bone from 14 prehistoric individuals. These individuals had originated from 36 individuals from the Gotland site in mainland Sweden and were chosen because of the large amounts of preserved mitochondrial DNA. To identify the gene mutations the polymorphism of the -13910T allele was amplified (Figure 5) (Malmström et al., 2010). The results showed that the frequency of the derived T allele is associated with the ability to digest unprocessed milk in adulthood (Malmström et al., 2010). Figure 5 shows that the PWC population had a significant -13910T frequency than the extent Swedish population (Malmström et al., 2010). This difference can be explained by genetic drift, which causes the changes in allele frequency of the same population separated by a large amount. There is also evidence of strong gene selection for the T gene that allows for lactose tolerance (Malmström et al., 2010). As seen from the analysis genetic drift and the switch to agricultural diet plays a role on lactose intolerance. Figure 5: Frequency of the -13910T allele in the lactase gene in three different populations, Swedish extant, Swedish Neolithic hunter-gatherer and the negative controls. Culture practices of the Scandinavian population played a role in the presentation of the -13910 allele. For instance, before the population practiced dairy farming, the allele frequency would have not been affected (Malmström et al., 2010). Also, the introduction of milk in Northern Europe would have heavily influenced the -13910T allele and this would have played a role in modern day culture, which is heavily influenced by dairy products. This could have resulted from the cross interaction of genes and culture (Malmström et al., 2010). Problems with DNA AnalysisThis section of the paper will focus on interpretation of ancient DNA and how we use it to understand human evolution. It will also be used to access the problems with ancient DNA and some of the challenges that are associated with it. Ancient DNA provides fundamental information on past disease pattern, geography and evolution. With every methodology such as aDNA there are always problems and thus it is important to explore the strengths and limitations. Interpretation of Microbial Ancient DNAOne of the most recent developments when using aDNA is that authentication cannot be done using only nucleotide sequences from ancient samples. Rollo et al., (1999) argues that sequence analysis should be the last step because there are large ranges of problems that are associated with this. For instance, contamination of specimens can occur through the introduction of modern microorganism to the ancient specimens. Although there is methodologies in place that limit the amount of contamination, it can still take part due to human error and other external factors. Rollo et al., (1999) statues that contamination can effect the ability to distinguish between DNA of ancient specimens and the DNA of microorganisms that have potentially colonized the remains. An example is seen through the reconstruction of the body of King Ramses II (1290-1224 BC). His body was so heavily colonized that when a sample was taken from his tissues 370 colonies were present and 89 different fungal species were isolated. Rollo et al., (1999) suggest that the solution would be to check the basis of the DNA sequence with present environmental characteristics from where the specimen originates. This methodology is important because it allows correct distinction of species that may be colonizing the ancient specimen. Extraction of specimens is important when preparing for DNA analysis. For instance in a study done by Rollo et al., (1999) a Neolithic herdsman hunter known as the Ice Man was excavated and microbial DNA was taken from the skin and the muscle to determine taphonomic history (Rollo et al., 1999). The specimen was swabbed with phenol, which removed any trace of ancient microbial colonization (Rollo et al., 1999). Without a viable sample, valuable information is lost and in turn also lost is the ability to understand disease origins and progression. Another issue faced by analysis to interpret ancient DNA, includes the ability to have DNA sequences that are conclusive for study. Problems with the DNA sequences can arise through post-mortem degradation of DNA from miscoding lesions or physical destruction of the molecule. The problems with the DNA sequence has led to major errors in studies that question the accuracy of disease origins and past population behaviour. Gilbert et al., (2005) proposes the nine criteria for authenticity (Figure 6). As seen from figure 6, the nine authenticity criteria involve: isolation from work areas, negative control extraction and amplification, appropriate molecular behaviour, reproducibility, cloning of products, independent replication, biochemical preservation, quantification and associated remains. The lack of compliance with the nine criteria can greatly affect the reliability and authenticity of the results. An issue with this list is that strict adherence does not mean full authenticity and can appear to be problematic. Ancient DNA studies are important when looking at modern diseases but the authenticity of molecular DNA strands is important when making inferences. This criterion is important to address when looking at the diseases because it plays a fundamental role in our understanding of disease origins and progression (Gilbert et al., 2005) Figure 6: Nine criteria for authenticity to determine the reliability of ancient DNA samples (Rollo et al., 1990: Box 1) These criteria have rarely been taken up completely in the field because there are instances there is lack of funding, and some believe there to be no contamination (Gilbert et al., 2005). As argued by Gilbert et al., (2005) it is important to implement these nine criteria because they assist in assessing ancient DNA studies more accurately. This is a growing field and thus new criteria and methodologies will be created to better handle ancient DNA specimens. Disease and Human EvolutionWhen analyzing disease using aDNA it is important to look at it through the context of human evolution. For instance, population genetic models can be used in an evolutionary aspect to predict the disease progression and susceptibility (Quintana-Murci et al., 2016). Ancient DNA samples in tangent with modern genomes gives the ability to reconstruct the genetic history of species that can shed light on disease. Firstly, the removal of deleterious mutations is important to understand the building blocks of human disease. Purifying selection is one of the most common forms of selection, which is the removal of select alleles that are highly associated with Mendelian disorders, or to maintain low population frequencies. The removal of deleterious alleles is fundamental when studying genetic disorders because it gives insight on population genetics and genetic drift. This can allow a human evolutionary perspective on DNA through its recreation of past populations while providing information on disease (Quintana-Murci et al., 2016). In continuation with human evolution and disease, rare variants provide valuable information on human disease. The recent study involving the 1000 genomes project revealed that there were a large number of rare variants (Quintana-Murci et al., 2016). Rare variants are genetic variants that alter gene function and play a role in Mendelian conditions. Although the direct contribution of rare variants are not finite, there presence in populations may cause underling early onset of disease and increase the susceptibility to common disease. Also, rare variants that are specific to a population cause more deleterious effects than common variants (Fig 7) (Quintana-Murci et al., 2016). Understanding rare variants in populations is important in aspects of human populations because it optimizes population sampling and identifying the rare variants that are disease causing (Quintana-Murci et al., 2016). This changed the way that DNA and ancient DNA are used to study common and rare disease. Figure 7: Demography in history that affects the proportion of deleterious variants on human population (Quintana-Murci et al., 2016: Figure 2) Figure 7 shows how demographic history affects deleterious and common variants differently. The proportion of variants is affected by the segregation on the populations, which is influenced by past demographics. The drawing illustrates the general demographic history of modern human populations, which include Africans, Europeans and French Canadians. The figure above does not represent change in population size but the deleterious variants in human population (Quintana-Murci et al., 2016). In terms of ancient DNA, the ability to recognize mutation through population genetics can assist in understanding human disease. Ancient DNA has made it possible to recognize the frequency of mutation in a population. Studies have shown that through the admixture of archaic variants and the genomes of modern humans, there have been improved adaptation and survival. Furthermore, deep sequencing is a new concept in aDNA studies that allows the sequencing of many different samples or populations changing the framework of aDNA analysis. Quintana-Murci et al., (2016) looked at a study of 230 human samples from West Eurasia that dates back to between 8500 and 2300 years ago. In the results of the study, they found 12 loci containing variants that changed through increasing positive selection. The variants included diet, genes encoding proteins involved in fatty acid metabolism, vitamin D, celiac disease and skin pigmentation (Quintana-Murci et al., 2016). Also, in this study it was found there was a positive selection for immunity related genes that are responsible for immune responses. This is important in aDNA studies because it can help to shed light on past human lifestyles and what selective events increased or decreased the frequency of certain alleles that are related to specific traits or disease (Quintana-Murci et al., 2016). Ancient DNA is fundamental when looking at the progression or incidence of disease because it can give insight into modern disease and rare or common diseases. Moreover, population genetics and human evolution are important components of ancient DNA studies because they are fundamental in studying disease. They provide insight into identification of alleles that are disease risk and the disease phenotypes (Quintana-Murci et al., 2016). Rare and deleterious variants are seen in both recent and ancient populations and how they have changed population frequencies and in turn effect survival (Quintana-Murci et al., 2016). More research needs to be conducted into looking at the entire human genome of current and past humans to ultimately determine the risk of disease on a molecular level. Ancient DNA and molecular phenotypic studies can be advantageous when looking at the relationship between genes, evolution and disease. ConclusionIn conclusion, studies of ancient DNA play a fundamental role in understanding disease patterns, disease progression and disease origins. As seen through the analysis of diseases such as malaria and tuberculosis, aDNA allows us to understand its origins and how it effected past populations. With this knowledge we are able to make inferences on modern populations and how disease will be affected on a population level. Looking at malaria has allowed for the understanding of the history of the disease through the analysis of Egyptian mummies and medieval individuals (Nerlich et al., 2008). Retrospective studies of malaria have allowed insight on future disease patterns. More specifically, the ability to identify Plasmodium falciparum on ancient specimen has allowed for dating and more insight on how it affects the human genome. Conversely, tuberculosis and Pott’s disease and identification through osteological evidence has allowed for a more in-depth understanding of the disease and its genome (Gernaey et al., 2001). This paper also looked at gene mutation in detail by analysing human evolution and lactose tolerance through the lactase gene. This has allowed for the identification of the T gene(-13910T) which is ultimately the cause of lactose intolerance (Myles et al., 2005). Gene mutation has allowed the understanding of culture and genetics to cross to provide an in-depth analysis on the disease. Gene mutation sheds light on how historical populations have affected modern day disease though the ability to consume milk and milk products. Lastly, this paper discussed the limitation of ancient DNA and its solutions. The nine criteria presented by Rollo et al., (1990) has allowed for authentication of studies to take place by having a test form contamination of DNA samples. Also, the study of human genomes through time can provide insight into disease. For instance, population genetics has allowed for the discovery of rare and deleterious variants (Quintana-Murci et al., 2016). In terms of ancient DNA, new deep sequencing methodologies allows analysis of multiple specimens and populations at once providing insight and comparison of disease (Quintana-Murci et al., 2016). Overall, aDNA is an important tool in anthropology and biology that can assist in disease diagnosis and predict the future of disease. Ancient DNA studies continue to grow and will allow for more in-depth accurate studies that will change the way modern diseases are viewed. ReferencesDonoghue, H. D., Spigelman, M., Greenblatt, C. L., Lev-Maor, G., Bar-Gal, G. K., Matheson, C., ... & Zink, A. R. (2004). Tuberculosis: from prehistory to Robert Koch, as revealed by ancient DNA.The Lancet infectious diseases,4(9), 584-592. Fletcher HA, Donoghue HD, Holton J, Pap I, Spigelman M. Widespread occurrence of Mycobacterium tuberculosis DNA from 18th–19th century Hungarians. Am J Phys Anthropol 2003; 120: 144–52. Gernaey, A. M., Minnikin, D. E., Copley, M. S., Dixon, R. A., Middleton, J. C., & Roberts, C. A. (2001). Mycolic acids and ancient DNA confirm an osteological diagnosis of tuberculosis.Tuberculosis,81(4), 259-265. Gilbert, M. T. P., Bandelt, H. J., Hofreiter, M., & Barnes, I. (2005). Assessing ancient DNA studies.Trends in ecology & evolution,20(10), 541-544. Haber, M., Mezzavilla, M., Xue, Y., & Tyler-Smith, C. (2016). Ancient DNA and the rewriting of human history: be sparing with Occam’s razor. Genome Biology, 17(1). Lalremruata, A., Ball, M., Bianucci, R., Welte, B., Nerlich, A. G., Kun, J. F., & Pusch, C. M. (2013). Molecular identification of falciparum malaria and human tuberculosis co-infections in mummies from the Fayum depression (Lower Egypt).PLoS One,8(4), e60307 Malmström, H., Linderholm, A., Lidén, K., Storå, J., Molnar, P., Holmlund, G., ... & Götherström, A. (2010). High frequency of lactose intolerance in a prehistoric hunter-gatherer population in northern Europe.BMC evolutionary biology,10(1), 89. Myles, S., Bouzekri, N., Haverfield, E., Cherkaoui, M., Dugoujon, J. M., & Ward, R. (2005). Genetic evidence in support of a shared Eurasian-North African dairying origin.Human genetics,117(1), 34-42. Nerlich, A. G., Schraut, B., Dittrich, S., Jelinek, T., & Zink, A. R. (2008). Plasmodium falciparum in ancient Egypt.Emerging infectious diseases,14(8), 1317. Quintana-Murci, L. (2016). Understanding rare and common diseases in the context of human evolution.Genome biology,17(1), 225. Rollo, F., & Marota, I. (1999). How microbial ancient DNA, found in association with human remains, can be interpreted.Philosophical Transactions of the Royal Society of London B: Biological Sciences,354(1379), 111-119. Suggested Reading from Inquiries Journal
Inquiries Journal provides undergraduate and graduate students around the world a platform for the wide dissemination of academic work over a range of core disciplines. Representing the work of students from hundreds of institutions around the globe, Inquiries Journal's large database of academic articles is completely free. Learn more | Blog | Submit Latest in Anthropology |